3.2.3. Inversion Input File
The inverse problem is solved using the executable program h3dtdinv_v2.exe. The lines of input file are as follows:
Line # |
Description |
Description |
|---|---|---|
1 |
path to tensor mesh file |
|
2 |
initial model |
|
3 |
reference model |
|
4 |
path to observations file |
|
5 |
sets time steps for the time-dependent problem |
|
6 |
topography |
|
7 |
currently ignored by program |
|
8 |
upper and lower bounds for recovered model |
|
9 |
additional cell weights |
|
10 |
cooling schedule for beta parameter |
|
11 |
weighting constants for smallness and smoothness constraints |
|
12 |
reference model update |
|
13 |
use SMOOTH_MOD or SMOOTH_MOD_DIFF |
|
14 |
stopping criteria for inversion |
|
15 |
parameters for Gauss-Newton iterations |
|
16 |
Solver |
choose Pardiso or MUMPS solver |
17 |
Memory Options |
store factorizations in RAM or write to disk |
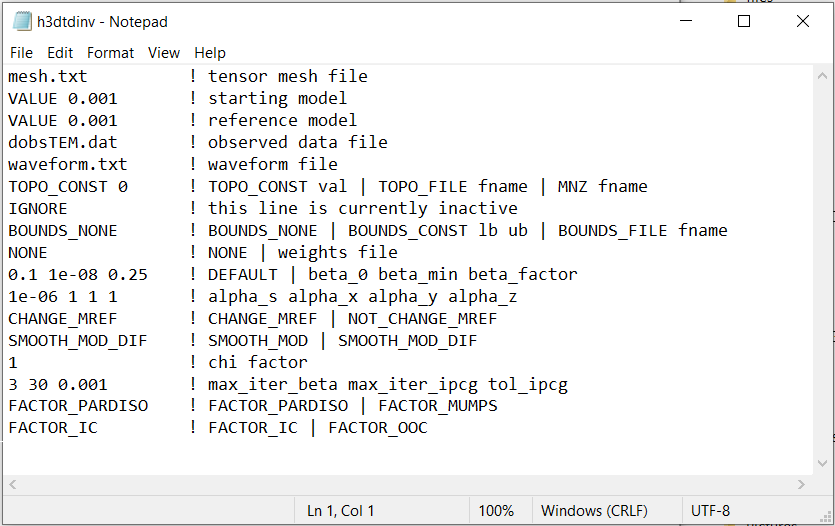
Fig. 3.3 Example input file for the inversion program (Download ).
3.2.3.1. Line Descriptions
Tensor Mesh: file path to the tensor mesh file
Initial Model: Defines the starting conductivity model for the inversion. There are two options:
FILE filepath: The user enters the flag FILE followed by the path to a conductivity model file
VALUE val: The user enters the flag VALUE followed by a value representing the conductivity of all cells lying below the surface topography
Reference Model: Defines the reference conductivity model for the inversion. There are two options:
FILE filepath: The user enters the flag FILE followed by the path to a conductivity model file
VALUE val: The user enters the flag VALUE followed by a value representing the conductivity of all cells lying below the surface topography
Observations File: Set the path to a observations file. The observations file defines the survey geometry, observed data and uncertainties.
Wave File: Set the path to a wave file. This file defines transmitter current and the time-stepping for the problem.
Active Topography Cells: Defines the active cells in the inversion. Choices are:
TOPO_CONST val: the flag ‘TOPO_CONST’ is followed by an elevation value for the topography
TOPO_FILE fname: the flag ‘TOPO_FILE’ is followed by the path to an active cells model
MNZ fname: flag ‘MNZ’ is followed by the path to an MNZ file; which contains an N x 3 array denoting the ijk indices of active cells.
IGNORE: Enter the flag IGNORE. This line is currently not used by the inversion code
Bounds: Bound constraints on the recovered model. There are 3 options:
BOUNDS_NONE: the flag BOUNDS_NONE is provided if there are no bounds on the recovered model
BOUNDS_CONST lb ub: the flag BOUNDS_CONST is entered followed by a lower and an upper bound value that will be applied to all cells; e.g. “BOUNDS_CONST 1E-6 0.1”
BOUNDS_FILE filepath: the flag BOUNDS_FILE is entered followd by the path to a bounds file
Weights: Here, the user specifies whether additional weights are supplied. If no additional weights are being supplied, enter the flag NONE. To apply weights, supply the path to a weights file.
beta_max beta_min beta_factor: Here, the user specifies protocols for the trade-off parameter (beta). beta_max is the initial value of beta, beta_min is the minimum allowable beta the program can use before quitting and beta_factor defines the factor by which beta is decreased at each iteration; example “1E4 10 0.2”. The user may also enter DEFAULT if they wish to have beta calculated automatically.
alpha_s alpha_x alpha_y alpha_z: Alpha parameters . Here, the user specifies the relative weighting between the smallness and smoothness component penalties on the recovered models.
Reference Model Update: Here, the user specifies whether the reference model is updated at each inversion step result. If so, enter CHANGE_MREF. If not, enter NOT_CHANGE_MREF.
Hard Constraints: if the flag SMOOTH_MOD is used, the reference model is not included in the smoothness terms of the model objective function; i.e. we preserve structures in the reference model but not their boundaries. If the flag “SMOOTH_MOD_DIF” is used, the reference model is included in the smallness and smoothness terms of the model objective function; i.e. we preserve the structures and boundaries defined in the reference model. For more, see the GIFtools cookbook .
Chi Factor: The chi factor defines the target misfit for the inversion. A chi factor of 1 means the target misfit is equal to the total number of data observations.
iter_per_beta max_iter_ipcg tol_ipcg: Here, iter_per_beta is the number of Gauss-Newton iterations performed for each beta value; see cooling schedule. max_iter_ipcg is the maximum number of iterations for the incomplete-preconditioned-conjugate gradient solve of the Gauss-Newton system, and tol_ipcg defines the tolerance (stopping criteria); see Gauss-Newton solve
Solver: Define the direct solver that will be used to factor and solve linear systems. Enter one of the following flags:
FACTOR_PARDISO: Factor and solve linear systems with Pardiso solver
FACTOR_MUMPS: Factor and solve linear systems with MUMPS solver
Memory Options: Enter one of the following flags:
FACTOR_IC: Store factorization in the computer’s RAM. This options is much faster but can only be used on problems of a reasonable size
FACTOR_OOC: Writes the factorizations of linear systems to disk. Slower but capable of solving much larger problems.